Eureka, we are back to spoilage yeasts. Today, I would like to introduce another spoilage yeast called Pichia kudriavzevii (anamorph Candida krusei). This yeast was first described by V.I. Kudryavtsev in 1960 as Issatchenkia orientalis but was classified as P. orientalis in 1964 and to P. kudriavzevii in 1965. P. kudriavzevii is a very abundant yeast in the environment and can be found in soil, fruits and various fermented beverages. So far, P. kudriavzevii is mainly associated with food spoilage to cause surface biofilms in low pH products [Kurtzman, 2011].
Where do I work?
Since this yeast is very abundant, P. kudriavzevii could be isolated from various sources: fruit juice, berries, sourdough, butter-like products, culture of Tanzanian fermented togw, African fermented cassava lafu, Ghanaian fermented cocoa bean heap, sake, champagne, ginger beer, tea beer, baker’s yeast, chicken egg, human heart blood, sputum, swine waste and human feces [Kurtzman, 2011; Chan 2012]. Well this does not sound very appetizing. And in fact, this strain is not really appetizing at all. Not only are various isolates from humans and animals of P. kudriavzevii available but many clinical cases support the idea that P. kudriavzevii (Candida krusei) is a species of clinical importance. In fact, Candida krusei is the 5th most common cause of candidemia [Kurtzman, 2011]. A kind of fungi infection which can affect immunocompromised patients (such as AIDS patients). Since we are already speaking of Candida, Candida albicans another species within the Candida family is one of the most pathogenic yeasts around and leads to Candidiasis. If you want to know how the symptoms of these medical conditions look like, feel free to google. But be warned, its nasty!
Back to the possible applications of P. kudriavzevii. As I already mentioned, P. kudriavzevii can be isolated from various sources and has been found in sourdough and further research needs to be performed to elucidate its possible function [Meroth 2003; De Vuyst, 2005]. Beside sourdough, P. kudriavzevii accounted for about 30% of the isolated yeasts in a cocoa bean heap fermentation and might be involved in the citrate assimilation during the fermentation [Daniel, 2009].
What about beer?
One existing association to beer is CBS strain 5148 which was isolated from a beer wort. Beside the “traditional” beer, one publication covers the isolation of P. kudriavzevii from tchapalo, a sorghum beer brewed in Côte d’Ivoire [N’guessan, 2011]. Whether P. kudriavzevii has any benefits for the production of beer like many other non-Saccharomyces strains has yet to be determined.
What is so special about me?
What is quite remarkable in my opinion is the fact that a draft genome assembly exists for a P. kudriavzevii strain (M12). Meaning the genome of this yeast has been sequenced and the information of the DNA sequences are known, published and available to everyone. Since DNA sequencing and the assembling process requires some financial investments as well as time (been there), there have to be certain reason(s) why such a research project is initiated and funded. And that’s where it gets interesting. Apparently, P. kudriavzevii contains some neat enzymes which make this strain very interesting in biotechnology for processes such as the production of bioethanol and phytases used to increase the uptake of phosphorus by plants (biofertilizer) [Chan, 2012]. Below is a simplified pathway of the Xylose pathway including the three enzymes found in P. kudriavzevii. Xylose by the way is a sugar molecule found in wood and not that many yeasts are capable of metabolizing this kind of sugar.

Xylulose-5-phosphate can then be input into the pentose phosphate pathway (PPP) to be converted into fructose 6-phosphate which can be passed to the glycolysis pathway. Under the right conditions, one might use P. kudriavzevii strain M12 to form alcohol from wood which is a very neat way of producing bioethanol.
Where can you find me?
Don’t know any commercial sources for P. kudriavzevii. This is for sure not a bad thing as I don’t want to have people playing around with possible infectious yeasts at home anyway…
Some biochemical stats about me for yeast ranchers
Below a summary of the biochemical properties of P. kudriavzevii. Data is summarized from Kurtzman et al (2011). Since this yeast can mainly ferment glucose, its spectra of sugars is very limited. For sure not suitable to ferment an entire batch of beer wort which contains a lot of maltose which cannot be utilized by P. kudriavzevii.
| Systematic name: | Pichia kudriavzevii (anamorph Candida krusei) | |
| Synonyms: | There are a lots of accepted synonyms for this yeasts. Just some examples: Saccharomyces krusei, Issatchenkia orientalis |
|
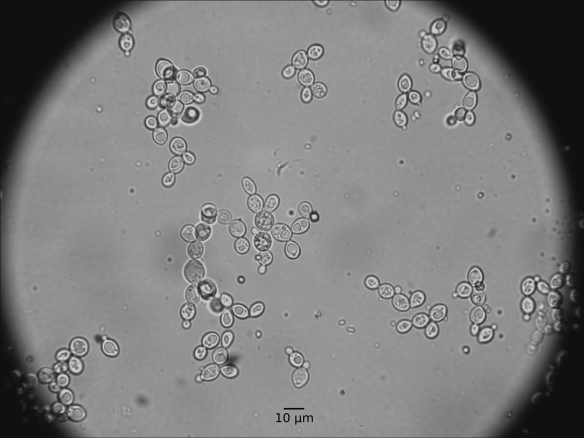
| Growth in malt extract: | Cell morphology: | Ovoid to elongate form, 1.3-6 µm x 3.3-14 µm |
| Clustering: | Occurring as single cells or in pairs |
|
| Pseudohyphae: | Moderate formation | |
| Pellicle formation: | Heavy, dry climbing pellicles are formed | |
| Growth in malt extract: | Colony morphology: | After 3 days: Butyrous and light-cream color |
| Fermentation: | Glucose: | Positive |
| Galactose: | Negative | |
| Sucrose: | Negative | |
| Maltose: | Negative | |
| Lactose: | Negative | |
| Raffinose: | Negative | |
| Trehalose: | Negative | |
That’s all about Pichia kudriavzevii so far. In summary, Pichia kudriavzevii seems to be involved in various beverage and food fermentations and is of clinical importance. Although the clinical importance mainly affects people with a suppressed immune system. It is not clear at this point if only a certain type of strain(s) (mainly the ones isolated from animals or humans) are potentially pathogenic or all the Pichia kudriavzevii strains.
Due to the potential pathogenic character as well as the limited range of sugars Pichia kudriavzevii can utilize, I don’t see any reasons to intentionally try this yeast for beer production. Furthermore, I do not support the idea to try the yeast for any beverages. Even if you could make beer out of wood…
Bibliography
- Chan GF, Gan HM, Ling HL, Rashid NA. (2012) Genome sequence of Pichia kudriavzevii M12, a potential producer of bioethanol and phytase. Eukaryot Cell, 11(10)
- Daniel HM, Vrancken G, Takrama JF, Camu N, De Vos P, De Vuyst L (2009) Yeast diversity of Ghanaian cocoa bean heap fermentations,FEMS Yeast Research, 9, 774–78
- De Vuyst L, Neysens P (2005) The sourdough microflora: biodiversity and metabolic interactions. Trends in Food Science & Technology, 16, 43–56
- Kurtzman CP, Fell JW, Boekhout T (2011) The Yeasts, a Taxonomic Study. Volume 1. Fifth edition. Elsevier (Link to sciencedirect)
- Meroth CB, Hammes WP, and Hertel C (2003) Identification and Population Dynamics of Yeasts in Sourdough Fermentation Processes by PCR-Denaturing Gradient Gel Electrophoresis. Applied and Environmental Microbiology, 69(12)
- N’guessan KF, Brou, K, Jacques N, Casaregola S, Dje, KM (2011) Identification of yeasts during alcoholic fermentation of tchapalo, a traditional sorghum beer from Côte d’Ivoire, Antonie van Leeuwenhoek, Vol99(4), 855-864